Molecular characterization and pathogenesis of gastrointestinal stromal tumor
Outline of the molecular pathogenesis of gastrointestinal stromal tumor (GIST)
GISTs are the most common mesenchymal tumors affecting the gastrointestinal tract (1). GISTs were formerly regarded as smooth muscle or neural neoplasms referred to as leiomyomas, leiomyosarcomas or schwannomas. However, identification of KIT mutations and high CD34 and c-KIT (CD117) positivity rates in these tumors led to the establishment of a new category of stromal tumors (2). The cellular origins of GISTs are thought to be interstitial cells of Cajal (ICCs), which are located in the myenteric plexus of the gastrointestinal tract, where they act as pacemaker cells for gastrointestinal motility. Subsequent studies showed that DOG1 (discovery on GIST1), also known as TMEM16A or ANO1, is a novel diagnostic marker of GISTs (3,4). Both DOG1 and KIT can serve as positive controls for immunohistochemical analysis in ICCs, though DOG1 is not expressed in KIT-positive mast cells (5). Protein kinase C θ (PKCθ) is specifically upregulated in GISTs as compared to other soft tissue tumors and, thus, it is also a useful diagnostic marker of GISTs (6).
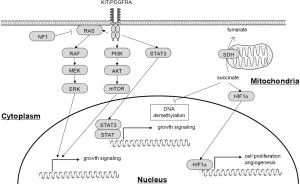
Activating mutations in the receptor tyrosine kinase gene KIT or platelet-derived growth factor receptor alpha (PDGFRA) play essential roles in the pathogenesis of GISTs through upregulation of downstream signaling pathways, including RAS/RAF/MAPK and PI3K/AKT/mTOR (Figure 1) (7). Mutations in RAS family genes and BRAF play a similar role, but are less frequently observed in GISTs (8). Succinate dehydrogenase (SDH)-deficient GISTs are characterized by wild-type KIT/PDGFRA and dysfunctional mutation or downregulation of members of the SDH heterotetramer (SDHA, SDHB, SDHC and SDHD). SDH deficiency and the resultant accumulation of succinate promote GIST development through different mechanisms than do oncogenic mutations, including upregulation of HIF1α and inhibition of DNA demethylation (Figure 1). Neurofibromin 1 (NF1) also acts as a tumor suppressor gene in GISTs, and patients with neurofibromatosis type I are known to be at high risk of developing multiple GISTs (9).
GISTs with no mutations in KIT, PDGFRA or RAS pathway genes or SDH-deficiency are referred as wild-type GISTs. They are characterized by overexpression of CALCRL/COL22A1, the tyrosine kinase NTRK2, the cyclin dependent kinase CDK6, and ERG, a member of the ETS-transcription factor family (10). A subset of wild-type GISTs exhibit mutations in TP53, MEN1 or MAX, and are characterized by a neural-committed phenotype and upregulation of the master endocrine regulator ASCL1 (11).
Chromosomal instability plays an important role in the development of many tumor types, and GISTs are characterized by various chromosomal abnormalities. For instance, losses of 14q and 22q frequently occur during the early stages of GIST development, and some of the chromosomal aberrations are associated with the clinical characteristics of GISTs (12). Epigenetic alterations, including aberrant DNA methylation and histone modification, have also been implicated in the development of GISTs (13,14). Recent studies have begun to shed light on the physiological and pathological importance of noncoding RNAs, and several noncoding RNAs are reportedly associated with the clinicopathological features of GISTs (15).
GISTs are rare tumors with an annual incidence of 10 to 20 per 1 million cases, but recent studies have shown that small GISTs may be occurring more frequently than previously documented. For instance, Agaimy et al. reported that microGISTs (less than 10 mm) are found in 22.5% autopsies performed in individuals older than 50 years (16). These lesions were located in the cardia, fundus, or proximal body of the stomach, but not in the antrum, duodenum, or remainder of the bowel. All tumors showed a histologically spindle cell morphology, and 57% of the tumors showed hyalinization and calcification. MicroGISTs were immunohistochemically positive for CD117, CD34, and vimentin, while KIT and PDGFRA mutations were found in 46% (11 of 24) and 4% (1 of 24) of these tumors, respectively (16). Kawanowa et al. investigated stomach specimens resected from 100 gastric cancer patients, and found a total of 50 microGISTs in 35 patients (17). All tumors were immunopositive for KIT or CD34 and negative for desmin. A large majority (45 of 50) of these tumors were located in the upper stomach, while only 8% (2 of 25) exhibited KIT mutation. In contrast to microGISTs, another study reported that KIT or PDGFRA mutations were detected in nearly all (12 of 13) small GISTs (less than 20 mm) (18). These results highlight the fact that although KIT/PDGFRA mutations are early events during GIST development, they are not sufficient for the progression of GISTs.
KIT mutations in GIST
KIT encodes the 145 kDa receptor tyrosine kinase c-KIT, which was identified as a normal cellular homolog of the feline sarcoma viral oncogene v-kit (19). KIT belongs to the type III receptor tyrosine kinase family, which includes PDGFRA, PDGFRB, macrophage colony stimulating factor receptor (CSF1R) and FL cytokine receptor (FLT3) (20). KIT is composed of an extracellular domain, juxtamembrane domain, tyrosine kinase domain I and tyrosine kinase domain II. KIT is maintained in an inactive form through auto-inhibition of the kinase domain (21).
Stem cell factor (SCF) is a KIT ligand, the binding of which promotes dimerization of the enzyme, ATP binding to the tyrosine kinase domain and auto phosphorylation of the tyrosine residue in the juxtamembrane domain (22). The SCF-KIT signal activates downstream pathways, including the MAP kinase cascade and PI3K/AKT pathway. The former leads to upregulation of such transcriptional factors as MYC, ELK, CREB and FOS, while the latter results in downregulation of cell cycle inhibitors and promotion of anti-apoptotic effects.
Approximately 70% to 80% of GISTs exhibit KIT mutations (23,24). The critical role of KIT mutation in GIST development has been well studied. For instance, the mutant forms of KIT protein harbor autonomous activity in the absence of ligand SCF binding (2), and a mutant Kit knock-in mouse model resembles familial GIST syndrome patients and shows diffuse ICC hyperplasia or GIST-like tumors (25,26). The mutant KIT activates multiple downstream signals, including MAPK, AKT, S6k, STAT1 and STAT3, in a SCF independent manner (27). The Kitv558Δ/+ mouse model shows that the PI3K/mTOR pathway is also upregulated in GISTs, and treatment with the mTOR inhibitor everolimus suppresses tumor proliferation (27). An ETS family member, ETV1, is regulated by active KIT, and cooperates with KIT to promote GIST growth. ETV1 is highly expressed in GISTs and acts as a transcriptional master regulator by binding to enhancer regions (28). ETV1 and KIT form a positive feedback loop to regulate target genes through stabilization of ETV1, and combination treatment with the KIT inhibitor imatinib and the MEK inhibitor MEK162 suppresses GIST growth in vivo and in vitro (29).
PDGFRA is another member of the receptor tyrosine kinase family and contributes to cell viability through ERK-dependent stabilization of ETV1 in KIT-mutant GISTs (30). Heat shock protein 90 (HSP90) is involved in the degradation of wild-type and mutant KIT (31), and a preclinical study showed that a HSP90 inhibitor promoted KIT degradation and suppressed GIST growth in vitro and in vivo (32). In a clinical trial, however, the response rate to IPI504, an ansamycin analogue HSP90 inhibitor, was low with a high toxicity rate (33). CDC37, a HSP90 cofactor, regulates KIT activation and expression and also interacts with oncogenic KIT (33).
Within GISTs, KIT mutations are found in several gene regions, including exons 8, 9, 11, 13, 14, 15, and 17. Exons 8 and 9 encode the extracellular domain, exon 11 encodes the juxtamembrane domain, and exons 13 and 17 encode the tyrosine kinase domain. Approximately 70% of GISTs exhibit mutations in exon 11, and 5% to 10% of GISTs show mutations in exon 9. Mutations in exon 11 disrupt auto-inhibition and lead to constitutive activation of KIT (34). Codons 557-558 in exon 11 are mutation hot spots, and deletions of W557 and/or K558 are associated with a metastatic phenotype (35) and poor post-operative recurrence-free survival (36). Another study showed that deletion-including codon 557/558 mutations are more strongly associated with larger tumor size, high mitotic count, high risk grade, and poor disease-free survival than other mutations in exon 11 (37). A small number of GISTs (6/427, 1.4%) show deletions in the boundary between intron 10 and exon 11, which could lead to loss of the normal splice acceptor site and p.K550_K558del mutation (23). GISTs with single nucleotide substitutions in exon 11 show indolent phenotype, lower mitotic activity, smaller tumor size, and favorable disease free survival (23,38). Within exon 11, tandem internal duplications occur mainly at the 3' end of the exon, and codons 576-579 are preferentially involved (23,39). Mutations in exon 9 are characterized by tandem duplication of six nucleotides at codons 502-503 (p.A502_Y503dup), and are associated with small bowel location, larger tumor size, older age (>60 years), female gender and spindle cell morphology (39).
Approximately 1% to 2% of KIT mutations are found in exons 13 and 17 (24,37,40). Most exon 13 mutations (e.g., c.1945A>G and c.1948G>A) result in p.K642E, which suppresses auto-inhibition of the juxtamembrane domain (41). About 70% of exon 17 mutations are c.2487T>A (p.N822K), while other infrequent mutations (p.N822Y, pN822K, p.N822H, p.D816F, p.D816Y, p.D820Y, p.D820V and p.Y823D) have also been identified (23,40). Exon 17 encodes the activation loop of the tyrosine kinase domain, and mutations in exon 17 are thought to be involved in maintenance of the constitutively active conformation (40). GISTs with mutations in exons 13 and 17 are associated with spindle cell morphology, and exon 13 mutations in particular correlate with the malignant potential of GISTs (40).
Mutations in exon 8 are rarely observed in GISTs, and in two cases with p.D419del mutation, one developed multiple peritoneal metastasis (42). Another study reported that, among three GISTs with exon 8 mutations (one case with p.D419del and two cases with heterozygous mutations of p.TYD417-419Y), all tumors were located at extragastric sites, and two cases showed distant metastasis (43). These reports suggest that mutations in exon 8 are potentially associated with the malignant phenotype of GISTs. Mutations in exon 14 are found as secondary mutations occurring after treatment with tyrosine kinase inhibitors (44,45). Mutations in exon 15 are rarely found in GISTs, and only c.2153C>G substitutions have been identified (46).
PDGFRA mutations in GIST
Approximately 10% to 15% of GISTs exhibit PDFRA mutations (47). These mutations are found in exon 12 (juxtamembrane domain), exon 14 (ATP biding domain), and exon 18 (activation loop), and cause constitutive PDGFRA activation in the absence of ligand binding, leading to downstream activation of signaling pathways. Like KIT mutations, PDGFRA mutations can activate a series of signal transduction molecules, including MAPK, AKT, STAT1 and STAT3 (47). HSP90 and a co-chaperone, CDC37, stabilize PDGFRA, and treatment with a HSP90 inhibitor represses AKT signaling (48). KIT and PDGFRA are close homologues, and their mutation occurs in a mutually exclusive manner. GISTs with PDGFRA mutations are characterized by gastric location, epithelioid morphology, and an indolent clinical course (49,50).
The most common PDGFRA mutation is p.D842V, which accounts for 60% to 65% of PDGFRA mutations in GISTs (approximately 5% of all GISTs) (23,37). This mutation is located in exon 18, a region encoding the second kinase domain, and is associated with extremely favorable disease-free survival as compared to other mutation types (37). Mutations in exon 14 are reportedly found in about 1% of all GISTs (51). The majority of exon 14 mutations are c.2125C>A or c.2125C>G missense mutations, which result in p.N659K, and c.2123A>T (p. N659Y) has also been reported (51). Mutations in exon 14 are associated with a gastric location, favorable clinical outcome and epithelioid morphology (51). Mutations in exon 12 are rarely observed (less than 1% of all GISTs) and include substitutions, small deletions and insertions (52). Locations and frequencies of KIT and PDGFRA mutations are summarized in Figure 2A.

Familial GIST
Familial GIST syndrome is characterized by germline mutation of KIT or PDGFRA, multiple GISTs, hyperpigmentation, mast cell tumors and ICC hyperplasia-associated dysphagia (53,54). KIT mutations observed in individuals with familial GIST include p.V559A, c.1756_1758delGAT and p.W557R in exon 11 (juxtamembrane domain) (55-57), deletion of one of two consecutive valine residues located between the transmembrane and tyrosine kinase domains (58), deletion of codon 419 in exon 8 (extracellular domain) (59), and D820Y substitution in exon 17 (53). A missense mutation (D846Y) in the exon 18 of PDGFRA has been also identified in familial GIST individuals (54). PDGFRA D846 is homologous to KIT D820, which is located within the tyrosine kinase domain. Most of the affected individuals develop multiple GISTs by middle age, and the tumors show histological features similar to sporadic GISTs, except for expansion of the myenteric plexus Cajal cell population (53). The ICC hyperplasia in familial GIST individuals represents non-neoplastic polyclonal proliferation, whereas GISTs in the same patients exhibit monoclonal proliferation (60). Mutations in familial GIST are summarized in Figure 2B.
SDH-deficient GIST
The most frequent molecular alteration in GISTs with wild-type KIT/PDGFRA is SDH deficiency. SDH consists of four subunits (SDHA, SDHB, SDHC, and SDHD), and is a component of the citric acid cycle and respiratory electron transfer chain (Figure 3) (61). SDH deficiency underlies Leigh syndrome, a neurodegenerative disorder caused by mitochondrial dysfunction, or several types of tumors, including paraganglioma, GIST, renal cell carcinoma and pituitary adenoma (62). SDH-deficient GISTs are immunohistochemically negative for SDHB due to its decreased expression or mutations in other SDH subunits that destabilize the SDH heterotetramer (63). Approximately 30% of SDHB-negative/SDH-deficient GISTs are also immunohistochemically negative for SDHA, the loss of which correlates generally with SDHA mutation (64). Patients with SDHA-positive GISTs are characterized by older age, female predominance, and a higher rate of liver metastasis than among those with SDHA-negative GISTs, although the mitosis rate, tumor size and clinical course are similar between SDHA-positive and -negative cases (64,65).

SDH deficiency results in the accumulation of succinate, which is a competitive inhibitor of ɑ-ketoglutarate-dependent dioxygenases, including the TET family of 5-methylcytosine hydroxylases (66). Members of the TET family are active DNA demethylases that convert 5-methylcytosine to 5-hydroxymethylcytosine, and inhibition of TET activities can lead to aberrant DNA methylation in GISTs. In fact, a genome-wide DNA methylation analysis of SDH-deficient GISTs revealed greater DNA hypermethylation than in GISTs with KIT mutation (67).
Accumulation of succinate is also involved in the stabilization of HIF1-ɑ, which controls oncogene transcription (68). Insulin-like growth factor 1 receptor (IGF1R) is overexpressed in KIT/PDGFR wild-type GISTs, and the expression is particularly elevated in SDH-deficient GISTs (69-71). The IGF family consists of two ligands (IGF1 and IGF2), two receptors (IGFR1 and IGFR1) and 6 IGF binding proteins (IGFBPs), and binding of IGF and IGFR activates downstream signals, including the MAPK and PI3K/AKT pathways (72). Inhibition of IGF1R induces apoptosis and represses AKT and MAPK signaling in GIST cells, which implicates the IGF signal in the development of SDH-deficient GISTs (73).
The Carney triad, Carney Stratakis syndrome, and several sporadic GISTs are included among the SDH-deficient GISTs (Figure 3) (1). Carney triad is characterized by gastric stromal sarcoma, paraganglioma, and pulmonary chondroma. It predominantly affects young females but has no heritability (74-76). Carney Stratakis syndrome is characterized by gastric GISTs and paragangliomas that exhibit mutation of the SDH subunits (77). This syndrome is inherited in an autosomal dominant manner, and some patients carry germline mutations in SDH family genes (64,65).
RAS signaling gene mutations in GIST
Mutations in RAS family genes and BRAF are found in a subset of GISTs. RAS proteins act as molecular switches that change between active GTP-bound and inactive GDP bound states. This switching mechanism is highly conserved among species, and conversion from the inactive GDP-bound form to the active GTP-bound form is mediated by guanine nucleotide exchange-factors (GEFs), while conversion back to the inactive form is mediated by GTPase-activating proteins (GAPs) (78). KRAS is frequently mutated in pancreatic, colorectal, and lung cancers, and most mutations occur at codon 12 or 13. The replacement of glycine at codon12 or 13 is thought to prevent inactivation by GAPs, which results in RAS activation in the absence of upstream stimulation (79). The BRAF V600E mutation is detected in malignant melanoma and thyroid and colorectal cancers (80-82). The mutant BRAF cooperates with Rac1b, AKT3 and other signal molecules to promote tumor cell viability and proliferation (83).
Miranda et al. detected KRAS mutations in 3 of 60 GISTs (5%) (8). In all three cases, the KRAS mutation was at codon 12 and/or 13 (G12D, G13D and G12A/G13D). The tumors carrying the G12D and G12A/G13D mutations showed deletions at exon 11 of KIT (Δ570-576 and Δ579), while the tumor with the G13D mutation exhibited PDGFRA mutation at exon 18 (D842V).
Multiple studies also identified the BRAF V600E mutation in GISTs with wild-type KIT/PDGFRA (84-86). Huss et.al. analyzed a cohort of 444 GISTs (272 KIT/PDGFRA-mutant and 172 wild type GISTs) and detected BRAF mutations in seven tumors (1.6% of all GISTs and 3.9% of wild-type GISTs) (87). Because BRAF mutation is found in small GISTs with diameters of 4 mm, it is considered to be one of the earliest events in the GIST development (88).
Other gene mutations in GIST
In addition to the mutations in well-known key driver genes, including KIT and PDGFRA, recent studies have revealed genetic alterations of other tumor-related genes in GISTs. For instance, EGFR mutations are found in 0.93% (3/323) of primary GISTs, and do not overlap with mutations in KIT, PDGFRA, KRAS or BRAF (89). EGFR mutations are associated with a stomach location, female gender and low recurrence rate. PIK3CA mutation (p.H1047L) has also been reported in a GIST case with KIT exon 11 deletion (84).
Analysis of 24 wild-type GISTs (without mutations in KIT/PDGFRA/RAS signal genes or SDH deficiency) identified 7 commonly mutated genes, ARID1B, ATR, FGFR1, LTK, SUFU, PARK2 and ZNF217 (90). Two of these tumors harbored FGFR1 gene fusions (FGFR1-HOOK3 and FGFR1-TACC1) and one exhibited ETV6-NTRK3 fusion. The ETV6-NTRK3 fusion transcript encodes the helix-loop-helix dimerization domain of ETV6 fused to the protein tyrosine kinase domain of NTRK3 (91), and the same fusion gene has been identified in breast carcinoma (92).
Alteration in protein phosphatase 2 regulatory subunit A alpha (PPP2R1A) causes dysfunction of protein phosphatase 2A (PP2A). Toda-Ishii et al. found PPP2R1A mutations in 17 of 94 (18%) GISTs, while a majority of the PPP2R1A mutant GISTs (16 of 17) harbored mutations in KIT, PDGFRA or RAS family genes and a remaining case showed SDH deficiency (93). BRCA1 and BRCA2 are well known tumor suppressor genes in breast and ovarian cancer, and a potential association between BRCA2 and GIST has been reported. An individual with a BRCA2 8642del3insC germline mutation developed prostate cancer, breast cancer and GIST (94).
Tumor suppressor genes in GIST
Neurofibromatosis type1 is an inheritable disease caused by bi-allelic loss of the NF1 gene (95). Neurofibromin contains a GAP-related domain (GRD) that is responsible for converting active Ras-GTP to inactive Ras-GDP, and negatively regulates RAS signaling. Individuals with NF1 mutations are at high risk of developing GISTs. NF1-associated GISTs are characterized by younger age at onset, location in the duodenum and small intestine, small size, tumor multiplicity and an indolent clinical course (9,96). Most NF1-associated GISTs are CD117-positive, have a spindle cell morphology, and generally show low mitotic rates. Hyperplastic foci (diffuse and focal) of CD117-positive ICCs are thought to be likely precursor lesions for GISTs, and precursors of NF1-associated GIST are often found around nerve plexuses. NF1-associated GISTs do not harbor KIT/PDGFRA mutations; instead, loss of NF1 leads to MAPK signal activation, while PI3K-AKT and JAK-STAT signals are less active than in common GISTs (97).
One recent study revealed that intragenic deletion of dystrophin (DMD) is a frequent event in metastatic GISTs (98). Dystrophin is expressed in sorted ICCs and inhibits GIST cell invasion, migration, anchorage independence and invadopodia formation, suggesting it plays a tumor suppressor and anti-metastatic role in GIST.
TP53 is the most frequently mutated gene in human malignancies. p53 acts as a tumor suppressor by mediating DNA repair, cell cycle arrest and apoptosis. Wild-type p53 is present at only low levels in normal cells due to its short half-life. TP53 mutant tumor cells are immunohistochemically positive for p53 because changes in its structure inhibit its ubiquitination and proteasomal degradation (99). Within GISTs, the rate of p53 positivity increases along with elevations in the mitotic index and tumor size (100). The p53 positivity is lower in gastric than intestinal GISTs, and is associated with epithelioid cell morphology, mucosal invasion, risk category and worse clinical outcomes (101). Murine double-minute 2 (MDM2) is an E3 ubiquitin ligase that negatively regulates p53 by mediating its ubiquitination and degradation (102). Induction of p53 through MDM2 inhibition exerts a moderate growth suppressive effect in TP53 wild-type GIST cells, suggesting p53 modulation may be an effective therapeutic strategy (103).
Chromosomal alterations in GIST
Chromosomal aberrations are prevalent among GISTs, with approximately 60% to 70% of all GISTs exhibiting alterations in chromosome 14, including loss of 14q and monosomy 14 (104,105). Loss of 14q is associated with gastric location, predominantly stable karyotypes, and favorable clinical outcomes (12). In addition, nearly half of GISTs show loss of 22q, while losses of 1p, 9p, 10q, 11p, 13q, 15q and 17p are also reported with lesser frequencies (12,106). Loss of 1p is associated with intestinal location, increased capacity for cytogenetic complexity and worse clinical outcomes, while loss of 22q is associated with increased capacity for cytogenetic complexity and poor disease-free survival (12). Losses of 9p, 11p and17p are also significantly associated with the GIST malignancy (104-107).
A number of functionally important genes are located in the regions frequently deleted in GISTs, including PARP2, APEX1, and NDRG2 at 14q11.2; SIVA at 14q32.33; MAX at 14q23.3; and NF2 at 22q12.2 (108). PARP2 suppresses genomic instability by regulating DNA repair and apoptosis (109). APEX1 also encodes a DNA repair enzyme implicated in the base excision pathway (110). NDRG2 is downregulated in various tumor types (111,112) and acts as a tumor suppressor by inhibiting tumor proliferation and promoting apoptosis (112,113). SIVA encodes a pro-apoptotic protein that binds to the tumor necrosis factor receptor CD27 (114). MAX encodes a basic helix-loop-helix leucine zipper transcription factor that interacts with MYC (115). Hemizygous or homozygous inactivating mutations of MAX are reported in 21% of all GISTs (17% of sporadic GISTs and 50% of sporadic and NF-1-associated GISTs) (115). Inactivation of MAX is also reported in microGISTs, suggesting its early onset during the development of GISTs (115). NF2 encodes the tumor suppressor protein merlin, which suppresses tumor cell growth by inhibiting the activities of RAS and RAC (108,116).
Gains and high level amplifications at 8q (including MYC) and 17q (including ERBB2) are significantly associated with metastatic GISTs, while those at 20q (including AIB1, AIB3, PTPN1 and MYBL2) are found in malignant primary and metastatic GISTs (105). AIB1, also referred to as nuclear receptor coactivator 3 (NCOA3), was first identified in a frequently amplified region in breast cancer (117). PTPN1 (also known as PTP1B) is involved in the regulation of cell growth, while MYBL2 is associated with cell cycle progression (118,119).
Epigenetic abnormalities in GIST
DNA methylation is an important mechanism for regulating gene expression, and hypermethylation of CpG islands is a major mechanism by which tumor suppressor genes are inactivated within tumor cells. Saito et al. analyzed a series of representative CpG islands and found methylation of MLH1, p73, p15, p16, CDH1 (E-cadherin), MGMT, MINT1 and MINT2 in GISTs, although the methylation status was not associated with KIT or PDGFRA mutations (120). They also concluded that 57% of GISTs exhibit hypermethylation of multiple CpG islands, which is referred as the CpG island methylator phenotype (120). Another study found that six genes (MGMT, p16, RASSF1A, CDH1, MLH1 and APC) are commonly methylated in GISTs and that methylation of CDH1 correlates with early recurrence and a poor prognosis in gastric GIST patients (13). p16 encodes a cyclin-dependent kinase inhibitor that negatively regulates G1/S-phase transition, while methylation and reduced p16 expression correlate with larger tumor size and poorer outcomes in GIST patients (121). A genome-wide DNA methylation analysis revealed that methylation of RASSF1A, REC8, and PAX3 are associated with the malignancy of GISTs (122).
Seventy to 80% of GISTs are immunohistochemically positive for the hematopoietic marker CD34 (123), and expression of CD34 is regulated through DNA methylation in gastric PDGFRA-mutant GISTs (124). Hypermethylation of PTEN is observed in GIST cells after long-term exposure to the tyrosine kinase inhibitor sunitinib, which suggests epigenetic silencing of PTEN may lead to drug-resistance in GISTs treated with tyrosine kinase inhibitors (125). Recent studies showed that microRNA (miRNA) genes are targets of aberrant DNA methylation in cancer, and we reported methylation-associated silencing of miR-34a and miR-335 in GIST cells (126).
DNA hypomethylation is associated with oncogene activation and chromosomal instability in various tumor types. ENDOGLIN/CD105 (ENG) is a transmembrane glycoprotein and auxiliary unit of the transforming growth factor-β (TGF-β) receptor encoded by ENG, which is overexpressed in KIT-positive GISTs (127). The elevated ENG expression is strongly associated with malignant and high-risk GISTs, and its overexpression is reportedly the result of DNA hypomethylation (127). About 45% of the human genome is composed of repetitive sequences, and methylation of long interspersed nuclear element-1 (LINE-1) is often used as a surrogate to evaluate global DNA hypomethylation in cancer. We reported that LINE-1 hypomethylation is strongly associated with clinical aggressiveness and DNA copy number aberrations in GISTs (128).
SETD2 is a histone methyltransferase that catalyzes methylation of histone H3 lysine 36 (H3K36), and trimethylation of H3K36 (H3K36me3) is a mark of active transcription (129). SETD2 mutations were recently identified in high-risk and metastatic GISTs (14). Loss of SETD2 is associated with reduced H3K36me3, DNA hypomethylated heterochromatin, and significantly worse outcomes in GIST patients, which suggests SETD2 is a novel GIST tumor suppressor (14).
Noncoding RNAs in GIST
Noncoding RNAs, including miRNAs and long noncoding RNAs (lncRNAs), play important roles in the development of various tumor types. miRNAs are small RNA molecules approximately 22 nt in length. Mature miRNAs are incorporated into RISC complexes and act to cleave complementary messenger RNA, or they repress translation by binding to the short complementary 3'-UTR region (130). Among their various functions, miRNAs are involved in cell proliferation, differentiation and apoptosis, and a number of miRNAs reportedly act as tumor suppressors or oncogenes (oncomir).
In GISTs, miRNA expression patterns are associated with tumor locations, risk classification and KIT/PDGRFRA mutation status (131,132). Because a large miRNA cluster is located in 14q32.31, loss of 14q is strongly associated with decreased expression of those miRNAs (131,132). Moreover, analysis using next generation sequencing identified a series of miRNAs differentially expressed in GISTs. These include miR-509-3p and miR-215-5p, expression of which is associated with cell type and risk grade (133). Another study showed that miR-133b is downregulated and its putative target gene, fascin-1, is overexpressed in high-risk GISTs (134). We showed that elevated expression of miR-196a is associated with high grade tumors and poor prognosis (15), while decreased expression of miR-186 correlates with post-operative recurrence (135). miRNAs also impact the drug sensitivities of GISTs, and overexpression of miR-125a-5p and miR-107 is associated with imatinib resistance (136). By contrast, miR-218 increases the sensitivity of GIST cells to imatinib by inhibiting the PI3K/AKT pathway (137).
Several studies have shown functional interactions between miRNAs and KIT in GISTs. For instance, expression of miR-221 and miR-222 correlates inversely with KIT expression in GISTs, suggesting these miRNAs may negatively regulate KIT expression (138). Other studies showed that members of the miR-17-92 and miR-221/222 clusters target KIT and ETV1 (139), and that miR-494 targets KIT (140). These results are indicative of the therapeutic potential of miRNAs for treatment of GISTs.
LncRNAs are generally defined as transcribed RNAs that do not have protein coding potential and are greater than 200 nt in length (141). LncRNAs exert their molecular effects by interacting with other cellular molecules, including DNA, protein and RNA, and through those interactions regulate various cancer-related pathways (142). Playing important roles in metastatic tumors, HOTAIR (HOX transcript antisense intergenic RNA) is one of the most extensively studied oncogenic lncRNAs (143,144). HOTAIR interacts with polycomb repressive complex 2 (PRC2) through its 5' terminal binding domain, and promotes H3K27me3-mediated gene silencing (145). We showed that overexpression of HOTAIR is associated with aggressiveness, and that HOTAIR knockdown suppressed the invasiveness of GIST cells (15). A more recent study showed that HOTAIR induces SUZ12-dependent hypermethylation of the protocadherin 10 (PCDH10) gene promoter in GIST cells, which further confirms the role of HOTAIR in GIST malignancy (146).
Conclusions
Molecular biological studies have greatly improved our understanding of the pathogenesis of GISTs, which has led to the successful use of receptor tyrosine kinase inhibitors for their treatment. In addition, recent advances in genomic and epigenomic analyses have enabled us to identify novel alterations that could be causally associated with GIST development. However, drug resistance due to additional mutations acquired during treatment remains a serious issue to overcome. Moreover, no specific treatments for wild-type GIST have yet been developed. It is anticipated that further molecular characterization of GISTs will contribute to the discovery of novel therapeutic targets and improved management of GISTs.
Acknowledgements
We thank Dr. William F. Goldman for editing the manuscript.
Footnote
Conflicts of Interest: The authors have no conflicts of interest to declare.
References
- Miettinen M, Lasota J. Histopathology of gastrointestinal stromal tumor. J Surg Oncol 2011;104:865-73. [Crossref] [PubMed]
- Hirota S, Isozaki K, Moriyama Y, et al. Gain-of-Function Mutations of c-kit in Human Gastrointestinal Stromal Tumors. Science 1998;279:577-80. [Crossref] [PubMed]
- West RB, Corless CL, Chen X, et al. The Novel Marker, DOG1, Is Expressed Ubiquitously in Gastrointestinal Stromal Tumors Irrespective of KIT or PDGFRA Mutation Status. Am J Pathol 2004;165:107-13. [Crossref] [PubMed]
- Espinosa I, Lee CH, Kim MK, et al. A novel monoclonal antibody against DOG1 is a sensitive and specific marker for gastrointestinal stromal tumors. Am J Surg Pathol 2008;32:210-8. [Crossref] [PubMed]
- Miettinen M, Wang ZF, Lasota J. DOG1 antibody in the differential diagnosis of gastrointestinal stromal tumors: a study of 1840 cases. Am J Surg Pathol 2009;33:1401-8. [Crossref] [PubMed]
- Duensing A, Joseph NE, Medeiros F, et al. Protein Kinase C θ (PKCθ) Expression and Constitutive Activation in Gastrointestinal Stromal Tumors (GISTs). Cancer Res 2004;64:5127-31. [Crossref] [PubMed]
- Duensing A, Medeiros F, McConarty B, et al. Mechanisms of oncogenic KIT signal transduction in primary gastrointestinal stromal tumors (GISTs). Oncogene 2004;23:3999-4006. [Crossref] [PubMed]
- Miranda C, Nucifora M, Molinari F, et al. KRAS and BRAF mutations predict primary resistance to imatinib in gastrointestinal stromal tumors. Clin Cancer Res 2012;18:1769-76. [Crossref] [PubMed]
- Andersson J, Sihto H, Meis-Kindblom JM, et al. NF1-associated gastrointestinal stromal tumors have unique clinical, phenotypic, and genotypic characteristics. Am J Surg Pathol 2005;29:1170-6. [Crossref] [PubMed]
- Nannini M, Astolfi A, Urbini M, et al. Integrated genomic study of quadruple-WT GIST (KIT/PDGFRA/SDH/RAS pathway wild-type GIST). BMC Cancer 2014;14:685. [Crossref] [PubMed]
- Pantaleo MA, Urbini M, Indio V, et al. Genome-Wide Analysis Identifies MEN1 and MAX Mutations and a Neuroendocrine-Like Molecular Heterogeneity in Quadruple WT GIST. Mol Cancer Res 2017;15:553-62. [Crossref] [PubMed]
- Gunawan B. An oncogenetic tree model in gastrointestinal stromal tumours (GISTs) identifies different pathways of cytogenetic evolution with prognostic implications. J Pathol 2007;211:463-70. [Crossref] [PubMed]
- House M. Tumor suppressor gene hypermethylation as a predictor of gastric stromal tumor behavior. J Gastrointest Surg 2003;7:1004-14. [Crossref] [PubMed]
- Huang KK, McPherson JR, Tay ST, et al. SETD2 histone modifier loss in aggressive GI stromal tumours. Gut 2016;65:1960-72. [Crossref] [PubMed]
- Niinuma T, Suzuki H, Nojima M, et al. Upregulation of miR-196a and HOTAIR drive malignant character in gastrointestinal stromal tumors. Cancer Res 2012;72:1126-36. [Crossref] [PubMed]
- Agaimy A, Wunsch PH, Hofstaedter F, et al. Minute gastric sclerosing stromal tumors (GIST tumorlets) are common in adults and frequently show c-KIT mutations. Am J Surg Pathol 2007;31:113-20. [Crossref] [PubMed]
- Kawanowa K, Sakuma Y, Sakurai S, et al. High incidence of microscopic gastrointestinal stromal tumors in the stomach. Hum Pathol 2006;37:1527-35. [Crossref] [PubMed]
- Anderson W, O'Sullivan B, Hughes F, et al. Microscopic gastrointestinal stromal tumours: a clinical and molecular study of 13 cases. Histopathology 2017;70:211-6. [Crossref] [PubMed]
- Besmer P, Murphy JE, George PC, et al. A new acute transforming feline retrovirus and relationship of its oncogene v-kit with the protein kinase gene family. Nature 1986;320:415-21. [Crossref] [PubMed]
- Hanks SK, Quinn AM, Hunter T. The protein kinase family: conserved features and deduced phylogeny of the catalytic domains. Science 1988;241:42. [Crossref] [PubMed]
- Mol CD, Dougan DR, Schneider TR, et al. Structural Basis for the Autoinhibition and STI-571 Inhibition of c-Kit Tyrosine Kinase. J Biol Chem 2004;279:31655-63. [Crossref] [PubMed]
- Lev S, Yarden Y, Givol D. Dimerization and activation of the kit receptor by monovalent and bivalent binding of the stem cell factor. J Biol Chem 1992;267:15970-7. [PubMed]
- Wozniak A, Rutkowski P, Piskorz A, et al. Prognostic value of KIT/PDGFRA mutations in gastrointestinal stromal tumours (GIST): Polish Clinical GIST Registry experience. Ann Oncol 2012;23:353-60. [Crossref] [PubMed]
- Debiec-Rychter M, Sciot R, Le Cesne A, et al. KIT mutations and dose selection for imatinib in patients with advanced gastrointestinal stromal tumours. Eur J Cancer 2006;42:1093-103. [Crossref] [PubMed]
- Rubin BP, Antonescu CR, Scott-Browne JP, et al. A Knock-In Mouse Model of Gastrointestinal Stromal Tumor Harboring Kit K641E. Cancer Res 2005;65:6631. [Crossref] [PubMed]
- Sommer G, Agosti V, Ehlers I, et al. Gastrointestinal stromal tumors in a mouse model by targeted mutation of the Kit receptor tyrosine kinase. Proc Natl Acad Sci U S A 2003;100:6706-11. [Crossref] [PubMed]
- Rossi F, Ehlers I, Agosti V, et al. Oncogenic Kit signaling and therapeutic intervention in a mouse model of gastrointestinal stromal tumor. Proc Natl Acad Sci U S A 2006;103:12843-8. [Crossref] [PubMed]
- Chi P, Chen Y, Zhang L, et al. ETV1 is a lineage survival factor that cooperates with KIT in gastrointestinal stromal tumours. Nature 2010;467:849-53. [Crossref] [PubMed]
- Ran L, Sirota I, Cao Z, et al. Combined inhibition of MAP kinase and KIT signaling synergistically destabilizes ETV1 and suppresses GIST tumor growth. Cancer Discov 2015;5:304-15. [Crossref] [PubMed]
- Hayashi Y, Bardsley MR, Toyomasu Y, et al. Platelet-Derived Growth Factor Receptor-alpha Regulates Proliferation of Gastrointestinal Stromal Tumor Cells With Mutations in KIT by Stabilizing ETV1. Gastroenterology 2015;149:420-32.e16. [Crossref] [PubMed]
- Fumo G, Akin C, Metcalfe DD, et al. 17-Allylamino-17-demethoxygeldanamycin (17-AAG) is effective in down-regulating mutated, constitutively activated KIT protein in human mast cells. Blood 2004;103:1078-84. [Crossref] [PubMed]
- Bauer S, Yu LK, Demetri GD, et al. Heat shock protein 90 inhibition in imatinib-resistant gastrointestinal stromal tumor. Cancer Res 2006;66:9153-61. [Crossref] [PubMed]
- Marino-Enriquez A, Ou WB, Cowley G, et al. Genome-wide functional screening identifies CDC37 as a crucial HSP90-cofactor for KIT oncogenic expression in gastrointestinal stromal tumors. Oncogene 2014;33:1872-6. [Crossref] [PubMed]
- Gajiwala KS, Wu JC, Christensen J, et al. KIT kinase mutants show unique mechanisms of drug resistance to imatinib and sunitinib in gastrointestinal stromal tumor patients. Proc Natl Acad Sci U S A 2009;106:1542-7. [Crossref] [PubMed]
- Wardelmann E, Losen I, Hans V, et al. Deletion of Trp-557 and Lys-558 in the juxtamembrane domain of the c-kit protooncogene is associated with metastatic behavior of gastrointestinal stromal tumors. Int J Cancer 2003;106:887-95. [Crossref] [PubMed]
- Martin-Broto J, Gutierrez A, Garcia-del-Muro X, et al. Prognostic time dependence of deletions affecting codons 557 and/or 558 of KIT gene for relapse-free survival (RFS) in localized GIST: a Spanish Group for Sarcoma Research (GEIS) Study. Ann Oncol 2010;21:1552-7. [Crossref] [PubMed]
- Wozniak A, Rutkowski P, Schoffski P, et al. Tumor genotype is an independent prognostic factor in primary gastrointestinal stromal tumors of gastric origin: a european multicenter analysis based on ConticaGIST. Clin Cancer Res 2014;20:6105-16. [Crossref] [PubMed]
- Steigen SE, Eide TJ, Wasag B, et al. Mutations in gastrointestinal stromal tumors – a population-based study from Northern Norway. APMIS 2007;115:289-98. [Crossref] [PubMed]
- Antonescu CR, Sommer G, Sarran L, et al. Association of Exon 9 Mutations with Nongastric Primary Site and Aggressive Behavior. Clin Cancer Res 2003;9:3329. [PubMed]
- Lasota J, Corless CL, Heinrich MC, et al. Clinicopathologic profile of gastrointestinal stromal tumors (GISTs) with primary KIT exon 13 or exon 17 mutations: a multicenter study on 54 cases. Mod Pathol 2008;21:476-84. [Crossref] [PubMed]
- Lux ML, Rubin BP, Biase TL, et al. KIT extracellular and kinase domain mutations in gastrointestinal stromal tumors. Am J Pathol 2000;156:791-5. [Crossref] [PubMed]
- Huss S, Kunstlinger H, Wardelmann E, et al. A subset of gastrointestinal stromal tumors previously regarded as wild-type tumors carries somatic activating mutations in KIT exon 8 (p.D419del). Mod Pathol 2013;26:1004-12. [Crossref] [PubMed]
- Ito T, Yamamura M, Hirai T, et al. Gastrointestinal stromal tumors with exon 8 c-kit gene mutation might occur at extragastric sites and have metastasis-prone nature. Int J Clin Exp Pathol 2014;7:8024-31. [PubMed]
- Nishida T, Kanda T, Nishitani A, et al. Secondary mutations in the kinase domain of the KIT gene are predominant in imatinib-resistant gastrointestinal stromal tumor. Cancer Sci 2008;99:799-804. [Crossref] [PubMed]
- Gao J, Tian Y, Li J, et al. Secondary mutations of c-KIT contribute to acquired resistance to imatinib and decrease efficacy of sunitinib in Chinese patients with gastrointestinal stromal tumors. Med Oncol 2013;30:522. [Crossref] [PubMed]
- Xu Z, Huo X, Tang C, et al. Frequent KIT mutations in human gastrointestinal stromal tumors. Sci Rep 2014;4:5907. [Crossref] [PubMed]
- Heinrich MC, Corless CL, Duensing A, et al. PDGFRA Activating Mutations in Gastrointestinal Stromal Tumors. Science 2003;299:708. [Crossref] [PubMed]
- Matei D, Satpathy M, Cao L, et al. The platelet-derived growth factor receptor alpha is destabilized by geldanamycins in cancer cells. J Biol Chem 2007;282:445-53. [Crossref] [PubMed]
- Lasota J, Dansonka-Mieszkowska A, Sobin LH, et al. A great majority of GISTs with PDGFRA mutations represent gastric tumors of low or no malignant potential. Lab Invest 2004;84:874-83. [Crossref] [PubMed]
- Sakurai S, Hasegawa T, Sakuma Y, et al. Myxoid epithelioid gastrointestinal stromal tumor (GIST) with mast cell infiltrations: A subtype of GIST with mutations of platelet-derived growth factor receptor alpha gene. Hum Pathol 2004;35:1223-30. [Crossref] [PubMed]
- Lasota J, Stachura J, Miettinen M. GISTs with PDGFRA exon 14 mutations represent subset of clinically favorable gastric tumors with epithelioid morphology. Lab Invest 2006;86:94-100. [Crossref] [PubMed]
- Corless CL, Schroeder A, Griffith D, et al. PDGFRA mutations in gastrointestinal stromal tumors: frequency, spectrum and in vitro sensitivity to imatinib. J Clin Oncol 2005;23:5357-64. [Crossref] [PubMed]
- Hirota S, Nishida T, Isozaki K, et al. Familial gastrointestinal stromal tumors associated with dysphagia and novel type germline mutation of KIT gene. Gastroenterology 2002;122:1493-9. [Crossref] [PubMed]
- Chompret A, Kannengiesser C, Barrois M, et al. PDGFRA germline mutation in a family with multiple cases of gastrointestinal stromal tumor. Gastroenterology 2004;126:318-21. [Crossref] [PubMed]
- Beghini A, Tibiletti MG, Roversi G, et al. Germline mutation in the juxtamembrane domain of the kit gene in a family with gastrointestinal stromal tumors and urticaria pigmentosa. Cancer 2001;92:657-62. [Crossref] [PubMed]
- Lasota J, Miettinen M. A new familial GIST identified. Am J Surg Pathol 2006;30:1342. [Crossref] [PubMed]
- Robson ME, Glogowski E, Sommer G, et al. Pleomorphic characteristics of a germ-line KIT mutation in a large kindred with gastrointestinal stromal tumors, hyperpigmentation, and dysphagia. Clin Cancer Res 2004;10:1250-4. [Crossref] [PubMed]
- Nishida T, Hirota S, Taniguchi M, et al. Familial gastrointestinal stromal tumours with germline mutation of the KIT gene. Nat Genet 1998;19:323-4. [Crossref] [PubMed]
- Hartmann K, Wardelmann E, Ma Y, et al. Novel germline mutation of KIT associated with familial gastrointestinal stromal tumors and mastocytosis. Gastroenterology 2005;129:1042-6. [Crossref] [PubMed]
- Chen H, Hirota S, Isozaki K, et al. Polyclonal nature of diffuse proliferation of interstitial cells of Cajal in patients with familial and multiple gastrointestinal stromal tumours. Gut 2002;51:793-6. [Crossref] [PubMed]
- Rutter J, Winge DR, Schiffman JD. Succinate dehydrogenase - Assembly, regulation and role in human disease. Mitochondrion 2010;10:393-401. [Crossref] [PubMed]
- Gill AJ, Lipton L, Taylor J, et al. Germline SDHC mutation presenting as recurrent SDH deficient GIST and renal carcinoma. Pathology 2013;45:689-91. [Crossref] [PubMed]
- Gaal J, Stratakis CA, Carney JA, et al. SDHB immunohistochemistry: a useful tool in the diagnosis of Carney-Stratakis and Carney triad gastrointestinal stromal tumors. Mod Pathol 2011;24:147-51. [Crossref] [PubMed]
- Dwight T, Benn DE, Clarkson A, et al. Loss of SDHA expression identifies SDHA mutations in succinate dehydrogenase-deficient gastrointestinal stromal tumors. Am J Surg Pathol 2013;37:226-33. [Crossref] [PubMed]
- Miettinen M, Killian JK, Wang ZF, et al. Immunohistochemical loss of succinate dehydrogenase subunit A (SDHA) in gastrointestinal stromal tumors (GISTs) signals SDHA germline mutation. Am J Surg Pathol 2013;37:234-40. [Crossref] [PubMed]
- Mason EF, Hornick JL. Succinate dehydrogenase deficiency is associated with decreased 5-hydroxymethylcytosine production in gastrointestinal stromal tumors: implications for mechanisms of tumorigenesis. Mod Pathol 2013;26:1492-7. [Crossref] [PubMed]
- Killian JK, Kim SY, Miettinen M, et al. Succinate dehydrogenase mutation underlies global epigenomic divergence in gastrointestinal stromal tumor. Cancer Discov 2013;3:648-57. [Crossref] [PubMed]
- Gill AJ. Succinate dehydrogenase (SDH) and mitochondrial driven neoplasia. Pathology 2012;44:285-92. [Crossref] [PubMed]
- Chou A, Chen J, Clarkson A, et al. Succinate dehydrogenase-deficient GISTs are characterized by IGF1R overexpression. Mod Pathol 2012;25:1307-13. [Crossref] [PubMed]
- Belinsky MG, Rink L, Flieder DB, et al. Overexpression of insulin-like growth factor 1 receptor and frequent mutational inactivation of SDHA in wild-type SDHB-negative gastrointestinal stromal tumors. Genes Chromosomes Cancer 2013;52:214-24. [Crossref] [PubMed]
- Lasota J, Wang Z, Kim SY, et al. Expression of the receptor for type i insulin-like growth factor (IGF1R) in gastrointestinal stromal tumors: an immunohistochemical study of 1078 cases with diagnostic and therapeutic implications. Am J Surg Pathol 2013;37:114-9. [Crossref] [PubMed]
- Yu H, Rohan T. Role of the insulin-like growth factor family in cancer development and progression. J Natl Cancer Inst 2000;92:1472-89. [Crossref] [PubMed]
- Tarn C, Rink L, Merkel E, et al. Insulin-like growth factor 1 receptor is a potential therapeutic target for gastrointestinal stromal tumors. Proc Natl Acad Sci U S A 2008;105:8387-92. [Crossref] [PubMed]
- Carney JA. Gastric stromal sarcoma, pulmonary chondroma, and extra-adrenal paraganglioma (Carney Triad): natural history, adrenocortical component, and possible familial occurrence. Mayo Clin Proc 1999;74:543-52. [Crossref] [PubMed]
- Zhang L, Smyrk TC, Young WF Jr, et al. Gastric stromal tumors in Carney triad are different clinically, pathologically, and behaviorally from sporadic gastric gastrointestinal stromal tumors: findings in 104 cases. Am J Surg Pathol 2010;34:53-64. [Crossref] [PubMed]
- Carney JA, Sheps SG, Go VL, et al. The triad of gastric leiomyosarcoma, functioning extra-adrenal paraganglioma and pulmonary chondroma. N Engl J Med 1977;296:1517-8. [Crossref] [PubMed]
- Carney JA, Stratakis CA. Familial paraganglioma and gastric stromal sarcoma: a new syndrome distinct from the Carney triad. Am J Med Genet 2002;108:132-9. [Crossref] [PubMed]
- Simanshu DK, Nissley DV, McCormick F. RAS Proteins and Their Regulators in Human Disease. Cell 2017;170:17-33. [Crossref] [PubMed]
- Scheffzek K, Ahmadian MR, Kabsch W, et al. The Ras-RasGAP complex: structural basis for GTPase activation and its loss in oncogenic Ras mutants. Science 1997;277:333-8. [Crossref] [PubMed]
- Cheung M, Sharma A, Madhunapantula SV, et al. Akt3 and Mutant V600 B-Raf Cooperate to Promote Early Melanoma Development. Cancer Res 2008;68:3429. [Crossref] [PubMed]
- Oliveira C, Velho S, Moutinho C, et al. KRAS and BRAF oncogenic mutations in MSS colorectal carcinoma progression. Oncogene 2007;26:158-63. [Crossref] [PubMed]
- Mitsutake N, Knauf JA, Mitsutake S, et al. Conditional BRAF V600E Expression Induces DNA Synthesis, Apoptosis, Dedifferentiation, and Chromosomal Instability in Thyroid PCCL3 Cells. Cancer Res 2005;65:2465. [Crossref] [PubMed]
- Matos P, Oliveira C, Velho S, et al. B-Raf(V600E) cooperates with alternative spliced Rac1b to sustain colorectal cancer cell survival. Gastroenterology 2008;135:899-906. [Crossref] [PubMed]
- Daniels M, Lurkin I, Pauli R, et al. Spectrum of KIT/PDGFRA/BRAF mutations and Phosphatidylinositol-3-Kinase pathway gene alterations in gastrointestinal stromal tumors (GIST). Cancer Lett 2011;312:43-54. [Crossref] [PubMed]
- Hostein I, Faur N, Primois C, et al. BRAF mutation status in gastrointestinal stromal tumors. Am J Clin Pathol 2010;133:141-8. [Crossref] [PubMed]
- Rossi S, Gasparotto D, Miceli R, et al. KIT, PDGFRA, and BRAF Mutational Spectrum Impacts on the Natural History of Imatinib-naive Localized GIST A Population-based Study. Am J Surg Pathol 2015;39:922-30. [Crossref] [PubMed]
- Huss S, Pasternack H, Ihle MA, et al. Clinicopathological and molecular features of a large cohort of gastrointestinal stromal tumors (GISTs) and review of the literature: BRAF mutations in KIT/PDGFRA wild-type GISTs are rare events. Hum Pathol 2017;62:206-14. [Crossref] [PubMed]
- Agaimy A, Terracciano LM, Dirnhofer S, et al. V600E BRAF mutations are alternative early molecular events in a subset of KIT/PDGFRA wild-type gastrointestinal stromal tumours. J Clin Pathol 2009;62:613-6. [Crossref] [PubMed]
- Shi SS, Wu N, He Y, et al. EGFR gene mutation in gastrointestinal stromal tumours. Histopathology 2017;71:553-61. [Crossref] [PubMed]
- Shi E, Chmielecki J, Tang CM, et al. FGFR1 and NTRK3 actionable alterations in "Wild-Type" gastrointestinal stromal tumors. J Transl Med 2016;14:339. [Crossref] [PubMed]
- Wai DH, Knezevich SR, Lucas T, et al. The ETV6-NTRK3 gene fusion encodes a chimeric protein tyrosine kinase that transforms NIH3T3 cells. Oncogene 2000;19:906-15. [Crossref] [PubMed]
- Tognon C, Knezevich SR, Huntsman D, et al. Expression of the ETV6-NTRK3 gene fusion as a primary event in human secretory breast carcinoma. Cancer Cell 2002;2:367-76. [Crossref] [PubMed]
- Toda-Ishii M, Akaike K, Suehara Y, et al. Clinicopathological effects of protein phosphatase 2, regulatory subunit A, alpha mutations in gastrointestinal stromal tumors. Mod Pathol 2016;29:1424-32. [Crossref] [PubMed]
- Waisbren J, Uthe R, Siziopikou K, et al. BRCA 1/2 gene mutation and gastrointestinal stromal tumours: a potential association. BMJ Case Rep 2015;2015. pii: bcr2014208830.
- Shen MH, Harper PS, Upadhyaya M. Molecular genetics of neurofibromatosis type 1 (NF1). J Med Genet 1996;33:2-17. [Crossref] [PubMed]
- Miettinen M, Fetsch JF, Sobin LH, et al. Gastrointestinal stromal tumors in patients with neurofibromatosis 1: a clinicopathologic and molecular genetic study of 45 cases. Am J Surg Pathol 2006;30:90-6. [Crossref] [PubMed]
- Maertens O, Prenen H, Debiec-Rychter M, et al. Molecular pathogenesis of multiple gastrointestinal stromal tumors in NF1 patients. Hum Mol Genet 2006;15:1015-23. [Crossref] [PubMed]
- Wang Y, Marino-Enriquez A, Bennett RR, et al. Dystrophin is a tumor suppressor in human cancers with myogenic programs. Nat Genet 2014;46:601-6. [Crossref] [PubMed]
- Baas IO, Mulder JW, Offerhaus GJ, et al. An evaluation of six antibodies for immunohistochemistry of mutant p53 gene product in archival colorectal neoplasms. J Pathol 1994;172:5-12. [Crossref] [PubMed]
- Al-Bozom IA. p53 expression in gastrointestinal stromal tumors. Pathol Int 2001;51:519-23. [Crossref] [PubMed]
- Feakins RM. The expression of p53 and bcl-2 in gastrointestinal stromal tumours is associated with anatomical site, and p53 expression is associated with grade and clinical outcome. Histopathology 2005;46:270-9. [Crossref] [PubMed]
- Moll UM, Petrenko O. The MDM2-p53 Interaction. Mol Cancer Res 2003;1:1001-8. [PubMed]
- Henze J, Muhlenberg T, Simon S, et al. p53 modulation as a therapeutic strategy in gastrointestinal stromal tumors. PLoS One 2012;7:e37776. [Crossref] [PubMed]
- Kim NG, Kim JJ, Ahn JY, et al. Putative chromosomal deletions on 9p, 9q and 22q occur preferentially in malignant gastrointestinal stromal tumors. Int J Cancer 2000;85:633-8. [Crossref] [PubMed]
- El-Rifai We, Sarlomo-Rikala M, Andersson LC, et al. DNA Sequence Copy Number Changes in Gastrointestinal Stromal Tumors: Tumor Progression and Prognostic Significance. Cancer Res 2000;60:3899. [PubMed]
- Wozniak A, Sciot R, Guillou L, et al. Array CGH analysis in primary gastrointestinal stromal tumors: cytogenetic profile correlates with anatomic site and tumor aggressiveness, irrespective of mutational status. Genes Chromosomes Cancer 2007;46:261-76. [Crossref] [PubMed]
- Gunawan B, Bergmann F, Höer J, et al. Biological and clinical significance of cytogenetic abnormalities in low-risk and high-risk gastrointestinal stromal tumors. Hum Pathol 2002;33:316-21. [Crossref] [PubMed]
- Assamaki R, Sarlomo-Rikala M, Lopez-Guerrero JA, et al. Array comparative genomic hybridization analysis of chromosomal imbalances and their target genes in gastrointestinal stromal tumors. Genes Chromosomes Cancer 2007;46:564-76. [Crossref] [PubMed]
- Fouquin A, Guirouilh-Barbat J, Lopez B, et al. PARP2 controls double-strand break repair pathway choice by limiting 53BP1 accumulation at DNA damage sites and promoting end-resection. Nucleic Acids Res 2017;45:12325-39. [Crossref] [PubMed]
- Madhusudan S, Smart F, Shrimpton P, et al. Isolation of a small molecule inhibitor of DNA base excision repair. Nucleic Acids Res 2005;33:4711-24. [Crossref] [PubMed]
- Kim YJ, Yoon SY, Kim JT, et al. NDRG2 expression decreases with tumor stages and regulates TCF/beta-catenin signaling in human colon carcinoma. Carcinogenesis 2009;30:598-605. [Crossref] [PubMed]
- Lorentzen A, Lewinsky RH, Bornholdt J, et al. Expression profile of the N-myc Downstream Regulated Gene 2 (NDRG2) in human cancers with focus on breast cancer. BMC Cancer 2011;11:14. [Crossref] [PubMed]
- Liu N, Wang L, Liu X, et al. Promoter methylation, mutation, and genomic deletion are involved in the decreased NDRG2 expression levels in several cancer cell lines. Biochem Biophys Res Commun 2007;358:164-9. [Crossref] [PubMed]
- Prasad KV, Ao Z, Yoon Y, et al. CD27, a member of the tumor necrosis factor receptor family, induces apoptosis and binds to Siva, a proapoptotic protein. Proc Natl Acad Sci U S A 1997;94:6346-51. [Crossref] [PubMed]
- Schaefer IM, Wang Y, Liang CW, et al. MAX inactivation is an early event in GIST development that regulates p16 and cell proliferation. Nat Commun 2017;8:14674. [Crossref] [PubMed]
- Morrison H, Sperka T, Manent J, et al. Merlin/Neurofibromatosis Type 2 Suppresses Growth by Inhibiting the Activation of Ras and Rac. Cancer Res 2007;67:520. [Crossref] [PubMed]
- Guan XY, Xu J, Anzick SL, et al. Hybrid selection of transcribed sequences from microdissected DNA: isolation of genes within amplified region at 20q11-q13.2 in breast cancer. Cancer Res 1996;56:3446-50. [PubMed]
- Tonks NK, Muthuswamy SK. A brake becomes an accelerator: PTP1B--a new therapeutic target for breast cancer. Cancer Cell 2007;11:214-6. [Crossref] [PubMed]
- Thorner AR, Hoadley KA, Parker JS, et al. In vitro and in vivo analysis of B-Myb in basal-like breast cancer. Oncogene 2009;28:742-51. [Crossref] [PubMed]
- Saito K, Sakurai S, Sano T, et al. Aberrant methylation status of known methylation-sensitive CpG islands in gastrointestinal stromal tumors without any correlation to the state of c-kit and PDGFRA gene mutations and their malignancy. Cancer Sci 2008;99:253-9. [Crossref] [PubMed]
- Mitomi H, Fukui N, Kishimoto I, et al. Role for p16(INK4a) in progression of gastrointestinal stromal tumors of the stomach: alteration of p16(INK4a) network members. Hum Pathol 2011;42:1505-13. [Crossref] [PubMed]
- Okamoto Y, Sawaki A, Ito S, et al. Aberrant DNA methylation associated with aggressiveness of gastrointestinal stromal tumour. Gut 2012;61:392-401. [Crossref] [PubMed]
- Miettinen M, Sobin LH, Sarlomo-Rikala M. Immunohistochemical Spectrum of GISTs at Different Sites and Their Differential Diagnosis with a Reference to CD117 (KIT). Mod Pathol 2000;13:1134-42. [Crossref] [PubMed]
- Bure I, Braun A, Kayser C, et al. The expression of hematopoietic progenitor cell antigen CD34 is regulated by DNA methylation in a site-dependent manner in gastrointestinal stromal tumours. Int J Cancer 2017;141:2296-304. [Crossref] [PubMed]
- Yang J, Ikezoe T, Nishioka C, et al. Long-term exposure of gastrointestinal stromal tumor cells to sunitinib induces epigenetic silencing of the PTEN gene. Int J Cancer 2012;130:959-66. [Crossref] [PubMed]
- Isosaka M, Niinuma T, Nojima M, et al. A Screen for Epigenetically Silenced microRNA Genes in Gastrointestinal Stromal Tumors. PLoS One 2015;10:e0133754. [Crossref] [PubMed]
- Gromova P, Rubin BP, Thys A, et al. ENDOGLIN/CD105 is expressed in KIT positive cells in the gut and in gastrointestinal stromal tumours. J Cell Mol Med 2012;16:306-17. [Crossref] [PubMed]
- Igarashi S, Suzuki H, Niinuma T, et al. A novel correlation between LINE-1 hypomethylation and the malignancy of gastrointestinal stromal tumors. Clin Cancer Res 2010;16:5114-23. [Crossref] [PubMed]
- Edmunds JW, Mahadevan LC, Clayton AL. Dynamic histone H3 methylation during gene induction: HYPB/Setd2 mediates all H3K36 trimethylation. EMBO J 2008;27:406-20. [Crossref] [PubMed]
- Bartel DP. MicroRNAs: Genomics, Biogenesis, Mechanism, and Function. Cell 2004;116:281-97. [Crossref] [PubMed]
- Choi HJ, Lee H, Kim H, et al. MicroRNA expression profile of gastrointestinal stromal tumors is distinguished by 14q loss and anatomic site. Int J Cancer 2010;126:1640-50. [PubMed]
- Haller F, von Heydebreck A, Zhang JD, et al. Localization- and mutation-dependent microRNA (miRNA) expression signatures in gastrointestinal stromal tumours (GISTs), with a cluster of co-expressed miRNAs located at 14q32.31. J Pathol 2010;220:71-86. [Crossref] [PubMed]
- Gyvyte U, Juzenas S, Salteniene V, et al. MiRNA profiling of gastrointestinal stromal tumors by next-generation sequencing. Oncotarget 2017;8:37225-38. [PubMed]
- Yamamoto H, Kohashi K, Fujita A, et al. Fascin-1 overexpression and miR-133b downregulation in the progression of gastrointestinal stromal tumor. Mod Pathol 2013;26:563-71. [Crossref] [PubMed]
- Niinuma T, Kai M, Kitajima H, et al. Downregulation of miR-186 is associated with metastatic recurrence of gastrointestinal stromal tumors. Oncol Lett 2017;14:5703-10. [PubMed]
- Akcakaya P, Caramuta S, Ahlen J, et al. microRNA expression signatures of gastrointestinal stromal tumours: associations with imatinib resistance and patient outcome. Br J Cancer 2014;111:2091-102. [Crossref] [PubMed]
- Fan R, Zhong J, Zheng S, et al. microRNA-218 increase the sensitivity of gastrointestinal stromal tumor to imatinib through PI3K/AKT pathway. Clin Exp Med 2015;15:137-44. [Crossref] [PubMed]
- Koelz M, Lense J, Wrba F, et al. Down-regulation of miR-221 and miR-222 correlates with pronounced Kit expression in gastrointestinal stromal tumors. Int J Oncol 2011;38:503-11. [Crossref] [PubMed]
- Gits CM, van Kuijk PF, Jonkers MB, et al. MiR-17-92 and miR-221/222 cluster members target KIT and ETV1 in human gastrointestinal stromal tumours. Br J Cancer 2013;109:1625-35. [Crossref] [PubMed]
- Kim WK, Park M, Kim YK, et al. MicroRNA-494 downregulates KIT and inhibits gastrointestinal stromal tumor cell proliferation. Clin Cancer Res 2011;17:7584-94. [Crossref] [PubMed]
- Iyer MK, Niknafs YS, Malik R, et al. The landscape of long noncoding RNAs in the human transcriptome. Nat Genet 2015;47:199-208. [Crossref] [PubMed]
- Schmitt AM, Chang HY. Long Noncoding RNAs in Cancer Pathways. Cancer Cell 2016;29:452-63. [Crossref] [PubMed]
- Liu XH, Liu ZL, Sun M, et al. The long non-coding RNA HOTAIR indicates a poor prognosis and promotes metastasis in non-small cell lung cancer. BMC Cancer 2013;13:464. [Crossref] [PubMed]
- Sorensen KP, Thomassen M, Tan Q, et al. Long non-coding RNA HOTAIR is an independent prognostic marker of metastasis in estrogen receptor-positive primary breast cancer. Breast Cancer Res Treat 2013;142:529-36. [Crossref] [PubMed]
- Bhan A, Mandal SS. LncRNA HOTAIR: A master regulator of chromatin dynamics and cancer. Biochim Biophys Acta 2015;1856:151-64.
- Lee NK, Lee JH, Kim WK, et al. Promoter methylation of PCDH10 by HOTAIR regulates the progression of gastrointestinal stromal tumors. Oncotarget 2016;7:75307-18. [PubMed]
Cite this article as: Niinuma T, Suzuki H, Sugai T. Molecular characterization and pathogenesis of gastrointestinal stromal tumor. Transl Gastroenterol Hepatol 2018;3:2.


